Tools
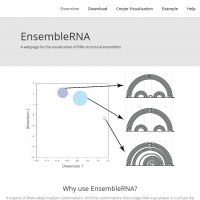
"A webpage for the visualization of RNA structural ensembles. A majority of RNAs adopt multiple conformations. All of the conformations that a single RNA may sample in a cell are the structural ensemble. RNA structure is most accurately visualized in this ensemble space. Visualization of the relationships between structures in an ensemble is key to understanding the effects of mutation or environment on RNA folding, stability and function."
"ERPIN (Easy RNA Profile IdentificatioN) takes as input an RNA sequence alignment and secondary structure annotation and will identify a wide variety of known RNA motifs (such as tRNAs, 5S rRNAs, SRP RNA, C/D box snoRNAs, hammerhead motifs, miRNAs and others) in your sequence(s) of interest. Also contains tool for drawing secondary structure of motifs."
"fRNAdb is a novel database service that hosts a large collection of non-coding transcripts including annotated/non-annotated sequences from the H-inv database,
NONCODE and RNAdb. A set of computational analyses have been performed on the included sequences. These analyses include RNA secondary structure motif discovery, EST support evaluation, cis-regulatory element search, protein homology search, etc."
Find download links for the Gencode V26 "Comprehensive gene annotation" set used in the construction of RNAStructuromeDB; region "CHR" and available for download as GTF or GFF3 file.
"Lncipedia.org is an integrated database of 146742 human annotated lncRNA transcripts obtained from different sources. In addition to basic transcript information and structure, several statistics are calculated for each entry in the database, such as secondary structure information, protein coding potential and microRNA binding sites."
"Long Noncoding RNA Database v2.0:The Reference Database For Functional Long Noncoding RNAs. Database that provides comprehensive annotations of eukaryotic long non-coding RNAs (lncRNAs)."
"The miRBase database is a searchable database of published miRNA sequences and annotation. Each entry in the miRBase Sequence database represents a predicted hairpin portion of a miRNA transcript (termed mir in the database), with information on the location and sequence of the mature miRNA sequence (termed miR). Both hairpin and mature sequences are available for searching and browsing, and entries can also be retrieved by name, keyword, references and annotation. All sequence and annotation data are also available for download."
"NRED integrates annotated expression data from various sources. Use this form to filter expression results data based on probe characteristics and/or the values of the expression data. If no experimental result set is selected, the form can also be used to search the probe table. All search fields are optional. For help and descriptions of the different fields, simply hover your mouse over the form labels."
"RBPmap is a computational tool that enables accurate prediction and mapping of RNA binding proteins (RBPs) binding sites on any RNA sequence or list of sequences of interest, provided by the users (as either sequences or genomic coordinates). RBPmap has been developed specifically for mapping RBPs in human, mouse and drosophila melanogaster genomes, though it supports mapping RBP binding sites in other organisms too."
"The Rfam database is a collection of RNA families, each represented by multiple sequence alignments, consensus secondary structures and covariance models (CMs)."
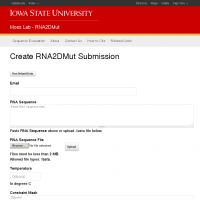
RNA2DMut will generate all possible point mutations of an input sequence and predicts structural information based on the Boltzmann 2D structural ensemble. RNA2DMut_Evaluate will calculate the same data, but for a set of user defined mutants.
"RNAcentral is a comprehensive database of non-coding RNA sequences that represents all types of ncRNA from a broad range of organisms"
This link will bring you to the download page for RNAz which "is a program for predicting structurally conserved and thermodynamically stable RNA secondary structures in multiple sequence alignments. It can be used in genome wide screens to detect functional RNA structures, as found in noncoding RNAs and cis-acting regulatory elements of mRNAs." The web-server version of this program can be found here: http://rna.tbi.univie.ac.at/
This is a link to the ViennaRNA Web Services. Here you can find RNA folding tools (including the RNAfold program - a version of which was used to construct the RNAStructuromeDB).