Frequently Asked Questions
Question:
Is there a way to automatically get outputs for a list of genes, one file per gene, using RNAStructuromeDB? Thanks!
Answer:
Yes! You can input a comma-separated list of gene names or ensemble IDs and download all results.
From the "Gene Search" tab change the filter for Gene Name (or Ensemble ID) to "Contains any word".
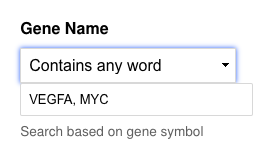
Here you can then input a list of gene names, and when you search all gene names matching those will be available for download at the bottom of the page (by clicking the red/orange CSV button).

Answered By:
Ryan J. Andrews
Category:
